Gene trapping involves technologies for marking genes by the random integration of DNA sequences containing a detectable reporter (marker). Gene trap technologies provide vectors that typically do not contain a promoter. Instead, gene trap vectors provide specific sequences that generate fusion RNA transcripts when inserted into a gene. This makes gene trapping very attractive for mammalian cells since these cells have complex genomic organization, including large introns and small exons. The trapped gene can then be identified by mRNA sequence. Using these technologies, it is possible to create libraries of cells in which the expression of the reporter gene provides measures of the activities of the trapped genes into which it is integrated. Subsequently, the gene in which the reporter gene has been inserted can be cloned using rapid amplification of cDNA (RACE) and have expression measured using fluorescence activated cell sorting (FACS).
However, there are limitations to these methods. Although an individual chemical compound can be screened for an effect on multiple genes simultaneously, the method entails the preparative scale use of FACS to enrich for cells that respond to a given modulator over multiple rounds of enrichment. As a consequence, prior methods are cost prohibitive and time consuming for use in high-throughput compound screens. Secondly, a library of cells containing trapped genes is divided into those that exhibit high levels of fluorescence and those that exhibit low levels of fluorescence i.e. essentially on or off. There is an inability to resolve changes in gene expression less than that required to cause a cell to change from on to off or off to on, and qualitative changes in gene expression are not identified.
Thus, there is a need in the field of drug screening to develop methodologies whereby a large number of compounds can be screened for their effects of the expression of genes and to develop methodologies whereby such effects can be qualitatively determined.
This invention provides a method for screening a large number of compounds for their inability to modulate the expression of genes. Specifically, this invention provides a method whereby trapped genes — whose activity is altered upon exposure to test compounds — are identified, cloned, and sequenced. Methodologies are provided that improve the accuracy, stability, and sensitivity of FACS screening, thus allowing multiplex screening of chemical compounds.
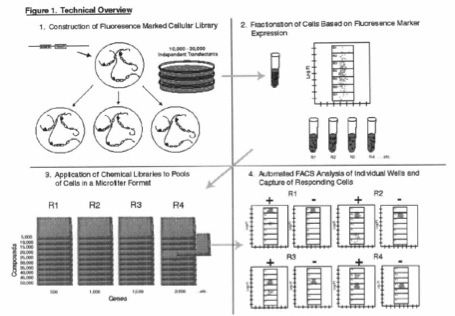
The method uses gene trap technology and comprises the steps of: transfecting a population of cells with a gene-trap vector, sorting cells according to their level of fluorescence, distributing sorted cells into pools, expanding the pools to obtain a sufficient number of cells representing each trapped gene to permit distinction of the effect of a test compound over controls, exposing the cells to the test compounds, and identifying compounds which alter the fluorescence distribution pattern of cells using FACS analysis.
Thus, this invention provides a method for high throughput multiplexed screening of test compounds for their effect on gene expression.