E3-Independent K867 monoubiquitination augments enzymatic activity of SETDB1
SETDB1 is an essential histone H3K9 methyltransferase and has been implicated in various physiological and pathological processes, such as development, endogenous retrovirus (ERV) silencing, neural disease, and cancer.
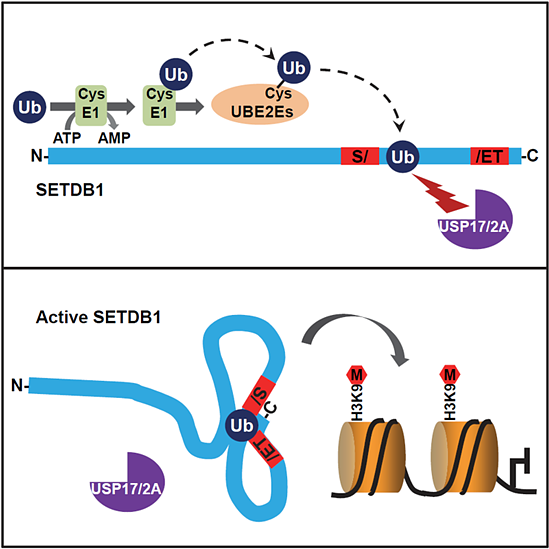
Different from most histone methyltransferases, the catalytic SET domain of SETDB1 is interrupted by a large 347-aa insertion conserved across different organisms but the function of this interposed sequence remains a mystery.
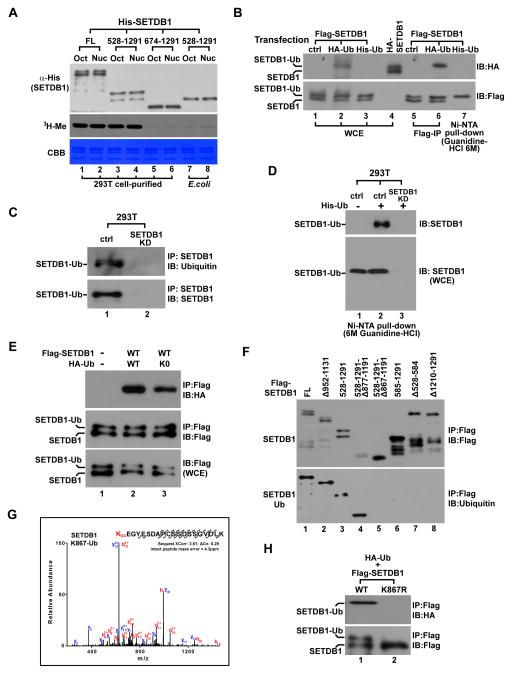
We recently demonstrated that SETDB1 is monoubiquitinated at the evolutionarily conserved lysine-867 (K867) within this insertion domain.
Intriguingly, K867 monoubiquitination is directly catalyzed by UBE2E family of E2 enzymes in an E3-independent manner while the conjugated ubiquitin (Ub) is protected from active deubiquitination.
The resulting constitutive lysine-867 monoubiquitination is essential for SETDB1’s enzymatic activity and endogenous retrovirus silencing in murine embryonic stem cells.
We also revealed that the canonical hydrophobic patch on the conjugated-Ub is critical for Ub protection and function. Together, our findings not only highlight a novel E3-independent mechanism for monoubiquitination, but also reveal molecular details underlying the regulation of SETDB1’s enzymatic activity.
We are now investigating the functional importance of lysine-867 monoubiquitination for SETDB1’s pro-oncogenic activities.
See the science
Connect with the Fang Lab
Department of Pharmacology and Therapeutics
Roswell Park Comprehensive Cancer Center
Elm and Carlton Streets
Buffalo, NY 14263